Micro Scale Thermophoresis
MicroScale Thermophoresis (MST) measures the motion of molecules along microscopic temperature gradients and detects changes in their hydration shell, charge or size.
When performing a MicroScale Thermophoresis (MST) experiment, a microscopic temperature gradient is induced by an infrared laser, and the directed movement of molecules is detected and quantified using either covalently attached dyes, fluorescent fusion proteins or intrinsic tryptophan fluorescence.
By combining the precision of fluorescence detection with the flexibility and sensitivity of thermophoresis, MST provides a flexible, robust and fast way to measure molecular interactions.
 The Monolith systems from NanoTemper provide an easy to use instrument line with minimal moving parts and therefore virtual maintenance free. Depending on the type the dynamic range is between 1 nM (1 pM) – mM and there is also a complete label free instrument. With low sample consumption (16 x 4 µl) per analysis, fast measurements (KD in 10 minutes) and rapid assay optimization, MST is a unique and straightforward platform.
The Monolith systems from NanoTemper provide an easy to use instrument line with minimal moving parts and therefore virtual maintenance free. Depending on the type the dynamic range is between 1 nM (1 pM) – mM and there is also a complete label free instrument. With low sample consumption (16 x 4 µl) per analysis, fast measurements (KD in 10 minutes) and rapid assay optimization, MST is a unique and straightforward platform.
MicroScale Thermophoresis (MST) is an easy, fast and precise way to quantify biomolecular interactions.
MST - Technology
Microscale Thermophoresis is an easy, fast and precise way to quantify biomolecular interactions. It measures the motion of molecules along microscopic temperature gradients and detects changes in their hydration shell, charge or size. It allows measuring a wide range of biomolecular interactions under close-to-native conditions: label-free, immobilization-free, in any buffer or complex bioliquid. NanoTemper's unique technology is ideal for basic research applications requiring flexibility in the experimental scale, as well as for pharmaceutical research applications including small molecules profiling.
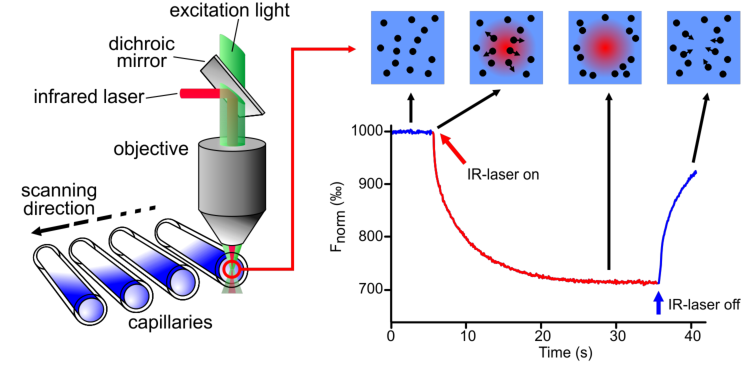
An infrared-laser is used to generate precise microscopic temperature gradients within thin glass capillaries that are filled with a sample in a buffer or bioliquid of choice. The fluorescence of molecules is used to monitor the motion of molecules along these temperature gradients. The fluorescence can be either intrinsic (e.g. tryptophan) or of an attached dye or fluorescent protein (e.g. GFP).
Sample preparation
To detect the thermophoretic mobility in a MST experiment, one of the interaction partners needs to be fluorescent. In case of the Monolith NT.LabelFree (and the LabelFree detector in the NT.Automated), the intrinsic tryptophan fluorescence of proteins can be exploited and no modification is required. For the other detectors, one of the interactants must be fluorescently labeled. Several coupling chemistries are available but the primary amine labeling with an N-hydroxysuccinimide (NHS) ester is one of the most commonly used for labeling proteins. NanoTemper Technologies offers labeling kits optimized for MST experiments. Alternatively, proteins can be expressed as fusion proteins (e.g. GFP), and biomolecules such as DNA and RNA oligos or peptides can be purchased from different companies carrying site-specific fluorophores.
Since MST measures the interaction at equilibrium, the interactants must be titrated against each other. The dilution series need to cover the baseline with fully unbound molecules and saturation of fully bound complex. This means that the dilution must be at least 20 fold below and above the dissociation constant. In the serial dilution, the labeled compound is kept constant and the binding partner is titrated.
After the making the dilution series the capillaries are filled with 4 µl of each of the samples.
The measurement
The capillaries are placed in the capillary tray and inserted in the instrument. The first scanning (the so-called “Cap Scan”) will locate the exact position of the capillaries and detect the overall fluorescence in the capillaries. To pinpoint the capillary position is essential to ensure correct focusing of the fluorescence detection and laser excitation with µm precision in the center of each capillary.
After the Cap Scan, the MST experiment can be conducted. In the figure the IR-laser is switched on after 5 seconds baseline measurement. Two effects happen when the laser is switched on: 1) a fast temperature jump (time scale ~ 1s) and the thermophoretic movement (timescale > 10s).
In a typical binding experiment, all 16 capillaries are measured within 10 min. For data analysis, the average thermophoretic value of each curve is plotted against the concentration of the binding partner resulting in a dose-response curve. The points are fitted and the KDof the interaction is determined.
MST experiments are performed to measure binding affinities, but the MST data and also the fluorescence signal recorded during the Cap Scan can provide information on sample property and homogeneity. These signals can be easily identified, thus assisting you to quickly optimize assay conditions:
- A lot of biomolecules stick to surfaces. Thus, they can also stick to the capillary wall making the solution inhomogeneous. This is easily detected from the shape of the “Cap Scan” and can be solved by employing a different capillary type or by using additives (detergents or BSA).
- The sample can be quenched in the solution resulting in different fluorescent signals per capillary. The nature of the quenching must be determined (sticking, aggregates, improper dilution, but also fluorescence quenching caused by a binding event) to facilitate appropriate assay optimization strategies or alternative data evaluation. NanoTemper Technologies assists you with short guidelines and protocols for straight-forward and simple assay optimization.
MST - Instruments
MicroScale Thermophoresis is an easy, fast and precise way to quantify biomolecular interactions. The MST technology provides numerous benefits:
- Optimize assays quickly: judge and improve sample quality immediately.
- Measure previously unmeasurable targets: work with very small amounts and sensitive samples.
- Benefit from close-to-native conditions: analyze in all bioliquids (cell lysate, serum) – immobilization-free and label-free.
- Do your research efficiently: enjoy perfect ease-of-use, purification-free measurements and get rid of maintenance downtimes.
- Work flexibly: K Ds for all molecule weights from ions to ribosomes and for pM to mM binding affinities.
The NT.115 Series measures biomolecular interactions via detection of fluorescent dyes or fluorescent fusion proteins (such as GFP) providing the following benefits:
- Immobilization-free affinity determination from 1 pM to mM.
- Broad application range
- Buffer independency: including serum or cell lysate
- Purification free: fluorescent fusion proteins
Watch the workflow of the NT.115 instruments at NanoTemper.
The Monolith NT.LabelFree measures MST via intrinsic tryptophan fluorescence [280 nm (ex) and 360 nm (em)], therefore providing a truly label-free reaction setup which is most beneficial for difficult samples such as membrane proteins. In addition, the experiment can be performed in any buffer. The NT.LabelFree is exceptionally sensitive for proteins binding to small molecules such as fragments, inhibitors or ions.
The Monolith NT.Automated is designed for high-throughput applications such as drug screening projects. The instrument comprises all benefits of the MST technology and is able to measure 96 samples in parallel. By integrating the instrument into a robot-compatible sample preparation platform, the overall handling time can be reduced to a minimum thus allowing for maximum automation. With the Monolith NT.Automated, NanoTemper Technologies offers a device which fulfills the high demands of pharmaceutical companies to screen and evaluate large libraries of drug candidates.
For a demonstration of the workflow of the NT.Automated, view the movies at NanoTemper.

MST - Applications
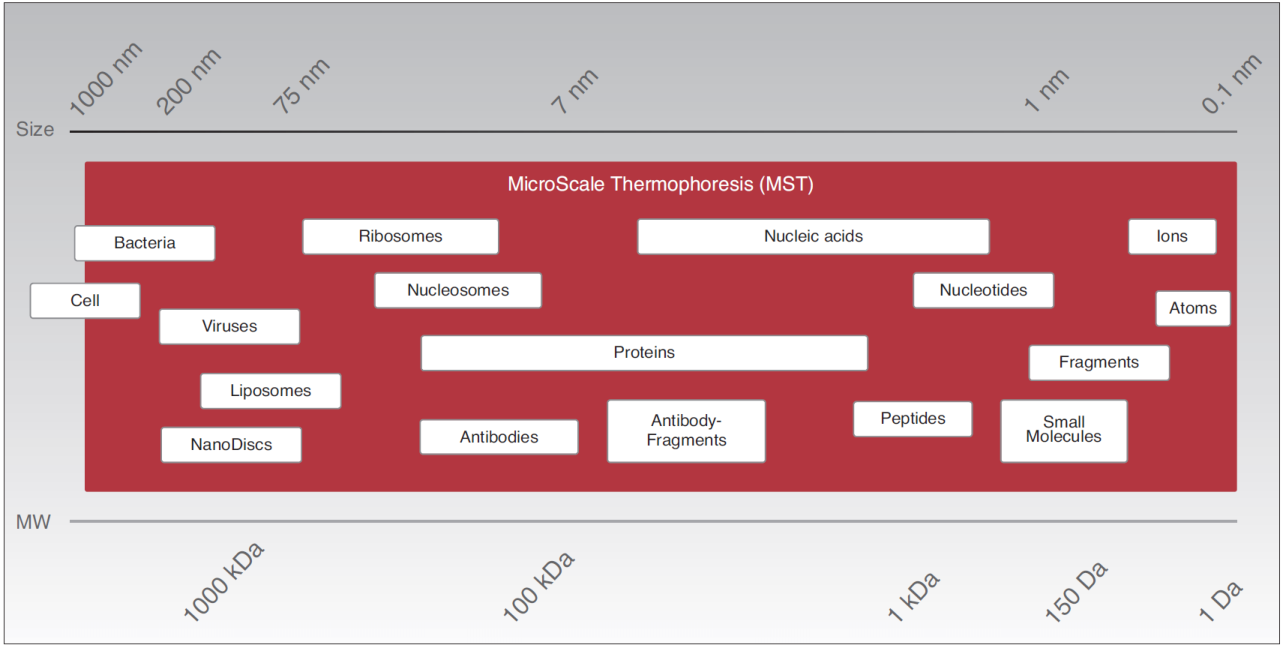
MicroScale Thermophoresis detects interactions between any kind of biomolecules thus providing a large application range, from ions and small molecules to high molecular weight and multi-protein complexes.
Thermophoresis, the movement of molecules in temperature gradients, is not only dependent on the size, but also on the charge and the hydration shell of the molecule of interest. Therefore, binding events can be detected even without an increase in size or mass upon complex formation.
Extensive research conducted at the Biophysics Department of the Ludwig-Maximilians-University Munich (LMU) (1) identified the solvation entropy and the hydration shell of molecules as the driving force. Any change of the hydration shell of biomolecules due to changes in their primary, secondary, tertiary and/or quaternary structure affects the thermophoretic movement and is used to determine binding affinities with high accuracy. The dependency on the energetic of the hydration shell allows for sensitivity unmatched by any other technology.
Since MST is performed free in solution without any surface immobilization, also bulky or sensitive molecule assemblies such as liposomes, nanodiscs or membrane proteins can be investigated.
Employing MST, the affinity between any kind of biomolecules but also other biophysical parameters such as stoichiometry, binding energetics, unfolding, can be determined.
- Biomolecular Interactions
- Affinity for 2, 3 or more interactants
- Label-free interactions [with MonoLith NT LabelFree]
- Purification-free interactions [with Monolith NT.115 Series instruments]
- Interactions in complex bio-liquids (serum, cell lysate) [with Monolith NT.115 Series instruments]
- Biophysical parameters (stoichiometry, binding energetics, unfolding, etc.
MST - Suppliers
NanoTemper Technologies GmbH
NanoTemper Technologies GmbH
Flößergasse 4
81369 München
Germany
Tel: +49 (0) 89 45 22 89 50
Fax: +49 (0) 89 45 22 895 60
website:
www.nanotempertech.com
e-mail :
Go to NanoTemper website for
application notes and publications.
References
| (1) | Seidel, S. A. I., P. M. Dijkman, W. A. Lea, et al. Microscale thermophoresis quantifies biomolecular interactions under previously challenging conditions. Methods 59: 301-315; (2013). Goto reference |